The table on the left is colored by triplet propensity, light-colored squares (like the GADV set) have high triplet propensity. The GADVESPLIT set are mostly higher than average, the GADV set really stands out. The correlation is too strong to be coincidence, so we can be confident that the earliest liforms had genes which popped into triplets with less energy input than is needed in modern organisms.
Why did ancient life-forms need to partition their nucleic acids into groups of three bases/basepairs? In modern life forms this partitioning is carried out by sophisticated enzymes as part of homologous recombination (exchange of genetic information for DNA repair or for sex). Perhaps in ancient lifeforms the biochemistry to perform these important activities was not so sophisticated, so the genome itself had to be structured so as to make these processes easier.
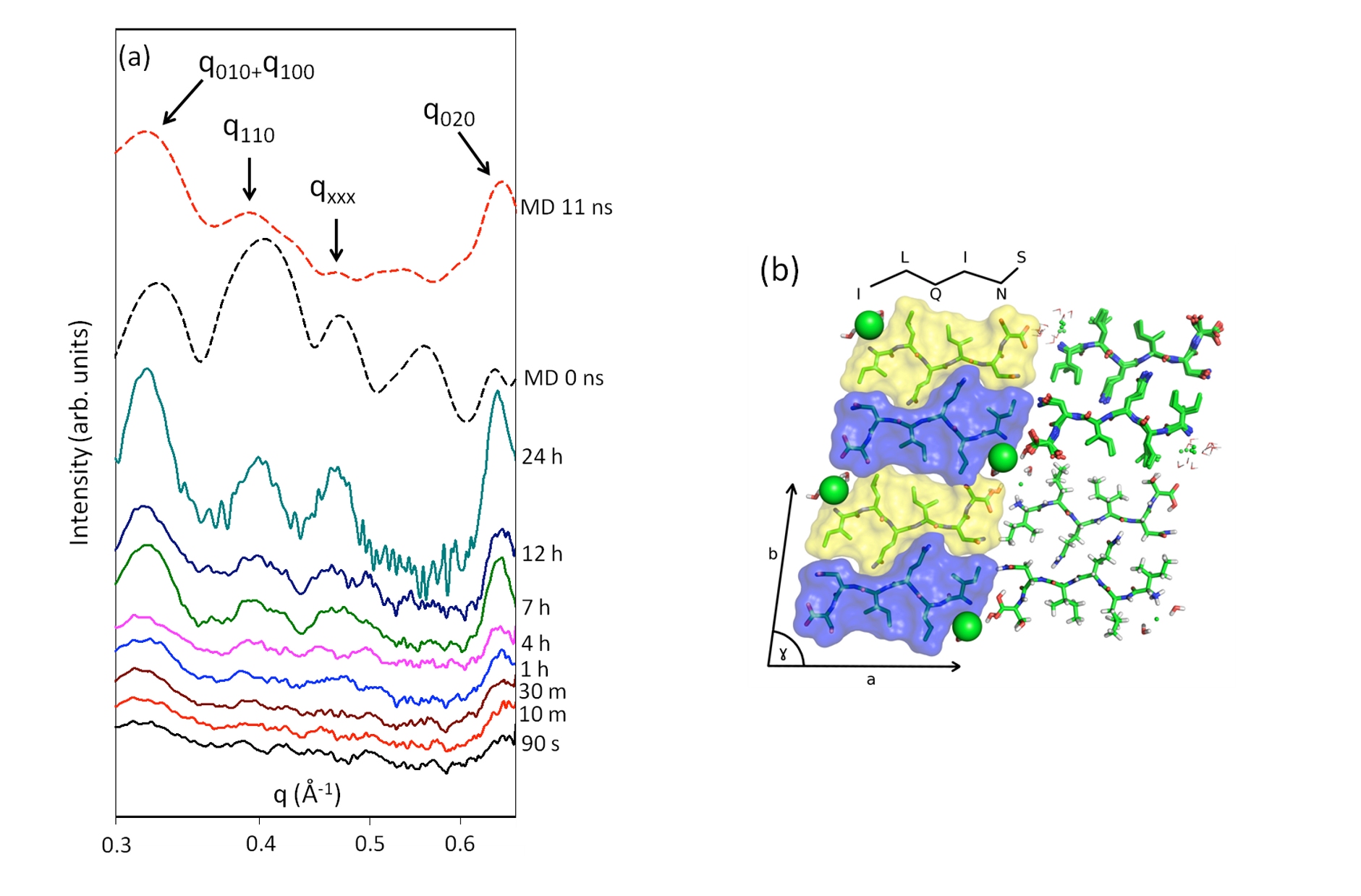 Amyloid Aggregation: Computer simulation can be used to `fill in the gaps' when experimental data is not completely explicit. The example figure shows some work to characterise the aggregation of ILQINS peptide (a digestion fragment of Hen's egg white lysozyme or "HEWL"). The spectra were measured using X-ray scattering by Nick Reynolds using the Australian Synchrotron. The structure is a guess made with the aid of MD simulations, which approximately reproduces the measured spectra. Hopefully as computer power becomes more freely available I will be able to begin using more systematic methods of structure determination, but for the moment this aspect of my work contains more art than science. Our latest paper on this topic is at Nature Comms.
Amyloid Aggregation: Computer simulation can be used to `fill in the gaps' when experimental data is not completely explicit. The example figure shows some work to characterise the aggregation of ILQINS peptide (a digestion fragment of Hen's egg white lysozyme or "HEWL"). The spectra were measured using X-ray scattering by Nick Reynolds using the Australian Synchrotron. The structure is a guess made with the aid of MD simulations, which approximately reproduces the measured spectra. Hopefully as computer power becomes more freely available I will be able to begin using more systematic methods of structure determination, but for the moment this aspect of my work contains more art than science. Our latest paper on this topic is at Nature Comms.
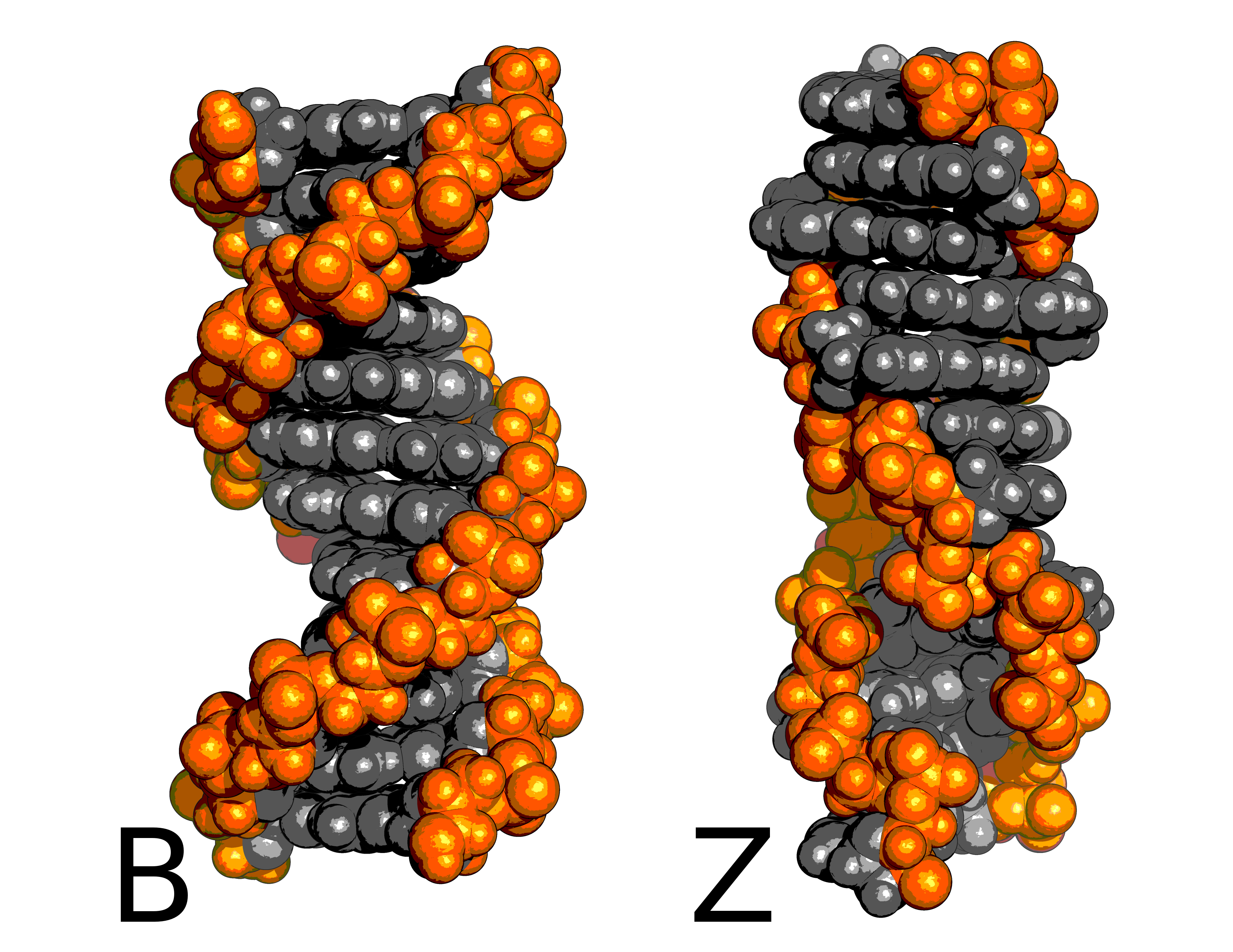 DNA Mechanics: DNA in high salt can switch helicity from right to left (B to Z). I am quite happy that Tanja and I were able to make the first accurate calculation of the concentration of salt needed to do this. I have some slides for a talk I gave, which might serve as an easy-reading introduction to the free energy method I used.
DNA Mechanics: DNA in high salt can switch helicity from right to left (B to Z). I am quite happy that Tanja and I were able to make the first accurate calculation of the concentration of salt needed to do this. I have some slides for a talk I gave, which might serve as an easy-reading introduction to the free energy method I used.
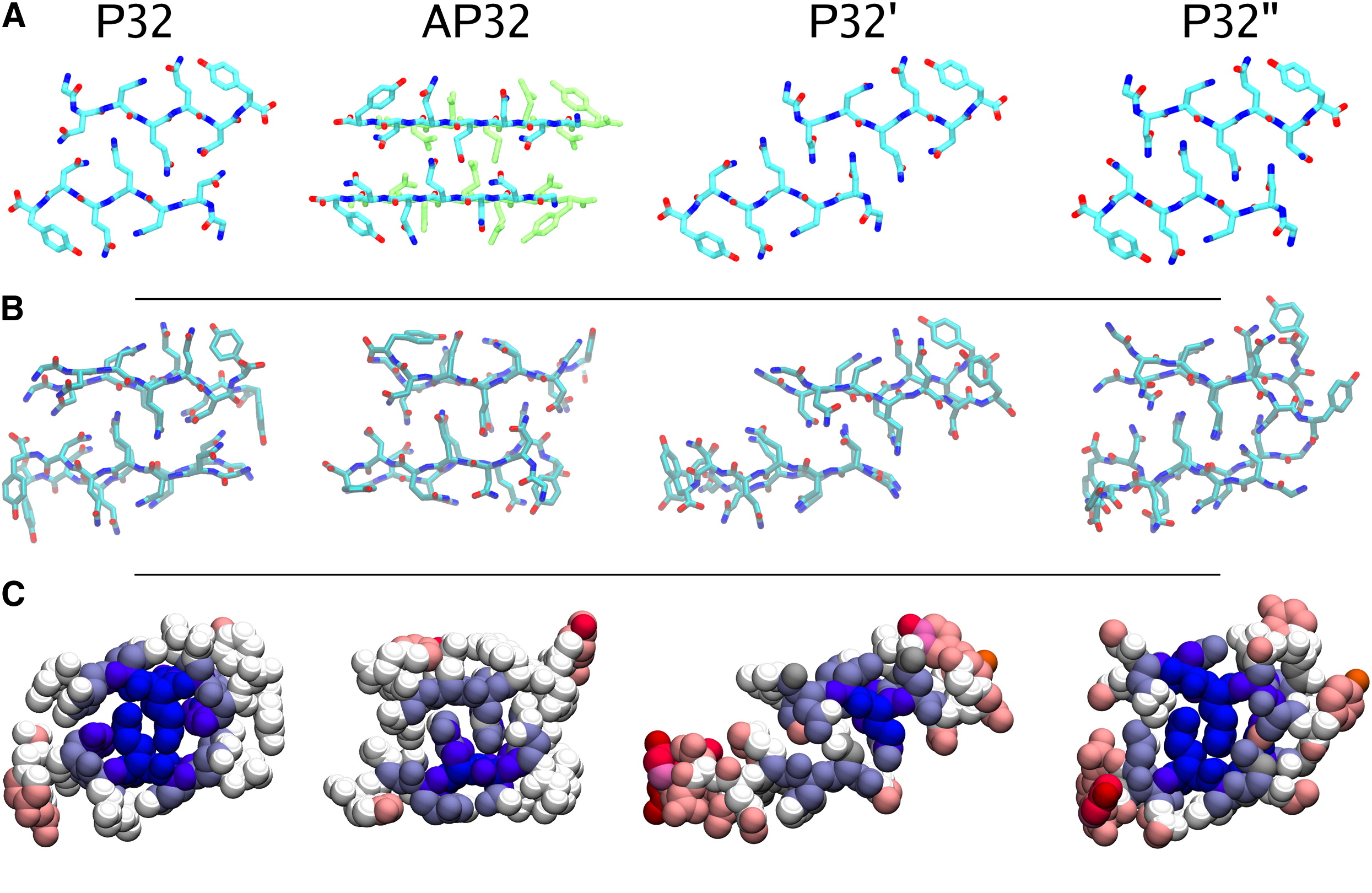
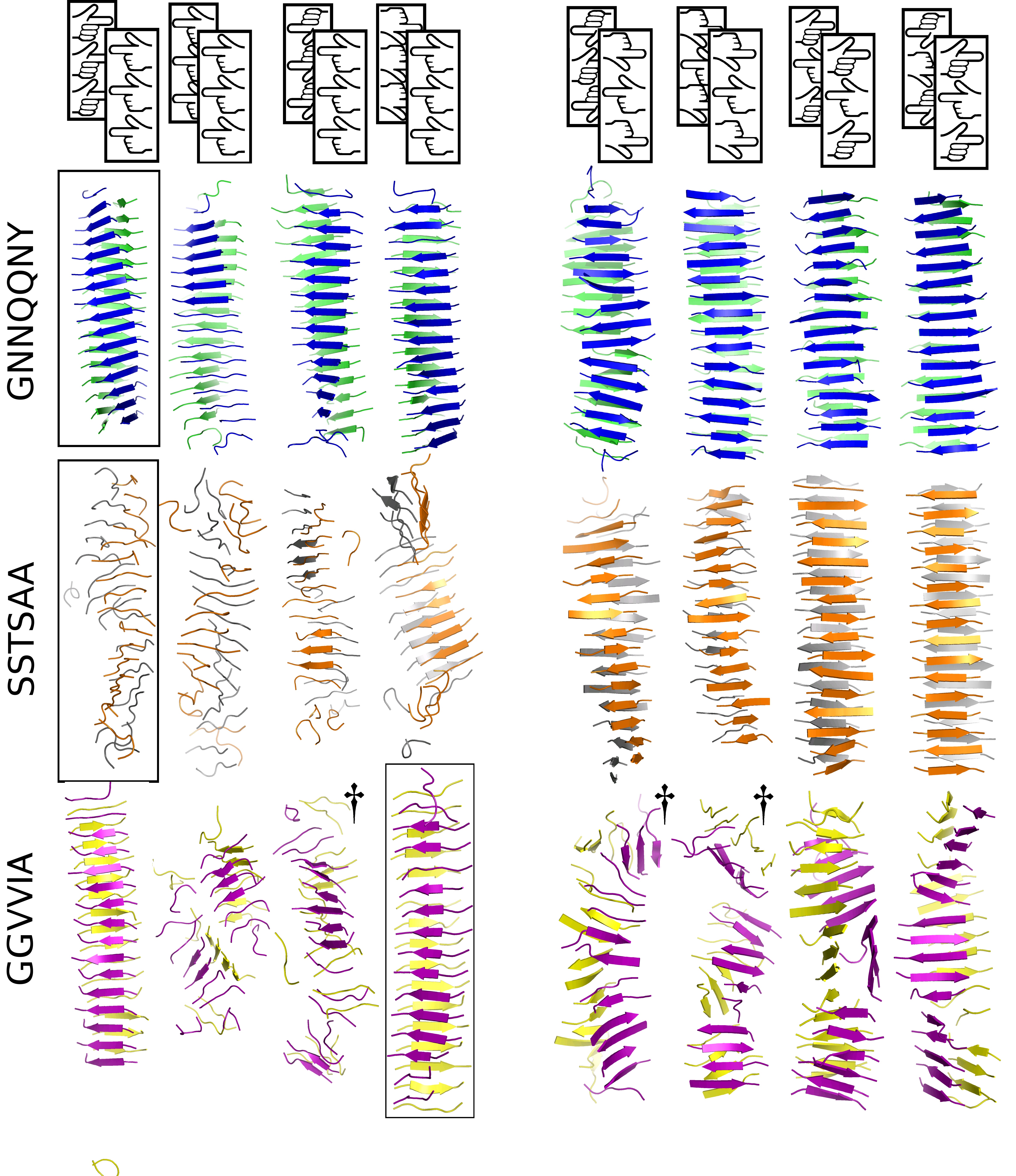
 Amyloid polymorphism: On the right is a journal cover from some of my PhD work. The image should hyperlink to the appropriate paper.
Amyloid polymorphism: On the right is a journal cover from some of my PhD work. The image should hyperlink to the appropriate paper.
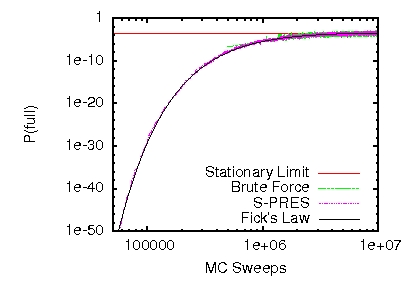 Rare Events: This figure illustrates the probability of a rare `jamming' event in a simple model of directed transport. Using S-PRES it was possible to measure events occuring at ridiculously low probabilities with negligible sampling error. Software now available for rare event calculation at freshs.org
Rare Events: This figure illustrates the probability of a rare `jamming' event in a simple model of directed transport. Using S-PRES it was possible to measure events occuring at ridiculously low probabilities with negligible sampling error. Software now available for rare event calculation at freshs.org